After 6 months, we were able to realize this project with all the hard work we put in. We were able to optimize an existent chromoprotein which allowed us to reach a bronze criterion. We were also able to produce a functional TAL, that’s able to recognize a specific sequence, which was able to answer the silver medal criterion. We were able to meet the requirements for a gold medal criterion with a successful model, the modeled was able to bring forth the rate of mortality by M.Tuberculosis and the survival rate using our detection kit.
Furthermore, we were able to further our project thanks to the integrated human practice by public engagement. That allowed us to optimize our kit and adapt it to the public needs. Those exchanges allowed our project to meet a silver criterion and the integration of the human practice meaning adapting our project to the answer of the surveys, permitted us to reach a Golden criterion.
PSR
We used the PSR (polymerase spiral reaction) method because it meets the criteria of our test. Indeed, it is a fast method, inexpensive and doesn’t require a great technical and material capacity. The important step of this achievement was to design specific pairs of primers that can amplify the DNA with a high specificity, but also sensitive to low concentrations of DNA.
Once the PSR method was well controlled, we were able to test our primers on a wide variety of bacterial strains and concluded that our primers had high specificity, which is essential for the diagnosis of M.Tuberculosis.
Sensitivity of this method is another important criterion, to be able to diagnose patients in an early stage of infection that would be undetectable by a microscope for example. After testing the sensitivity of this method with different DNA concentration, we noticed that our primers have a sensitivity equivalent to a PCR test. We can see 10X more sensitive for M.Smegmatis, and the limit of detection is up to 100 copies of genomic DNA per microliter, which is a good limit of detection for a testing based in molecular biology. Click here for more.
TAL
TALs proteins are naturally produced in genus Xanthomonas bacteria that are able to infect plants by binding itself specifically to the promoter. In our project, we designed our TALs protein to recognize a sequence of choice. We were able to produce it in E. coli, we successfully produced, and purified a functional TAL2 (BBa_K2915275). Then tested it on a gel shift, which confirms that our TAL2 is functional and specific to a specific sequence chosen.
We’re able to choose our sequence of choice by basing on the DNA fragment amplify (5’ CCGCTG 3’). The sequence that was chosen are found on rpoB gene and katG gene of M. Tuberculosis, dependent on which primers are used to amplify the DNA. There’s a beginning of sensitivity detected in between the DNA amplify and TAL2 with a ratio of 1:30 (see more). This part BBa_K2915275 is our validated part for a Silver medal criteria.
We successfully characterized an existing iGEM biobrick (BBa_K1033929). We determined its extinction coefficient and its thermal stability. See here for more details
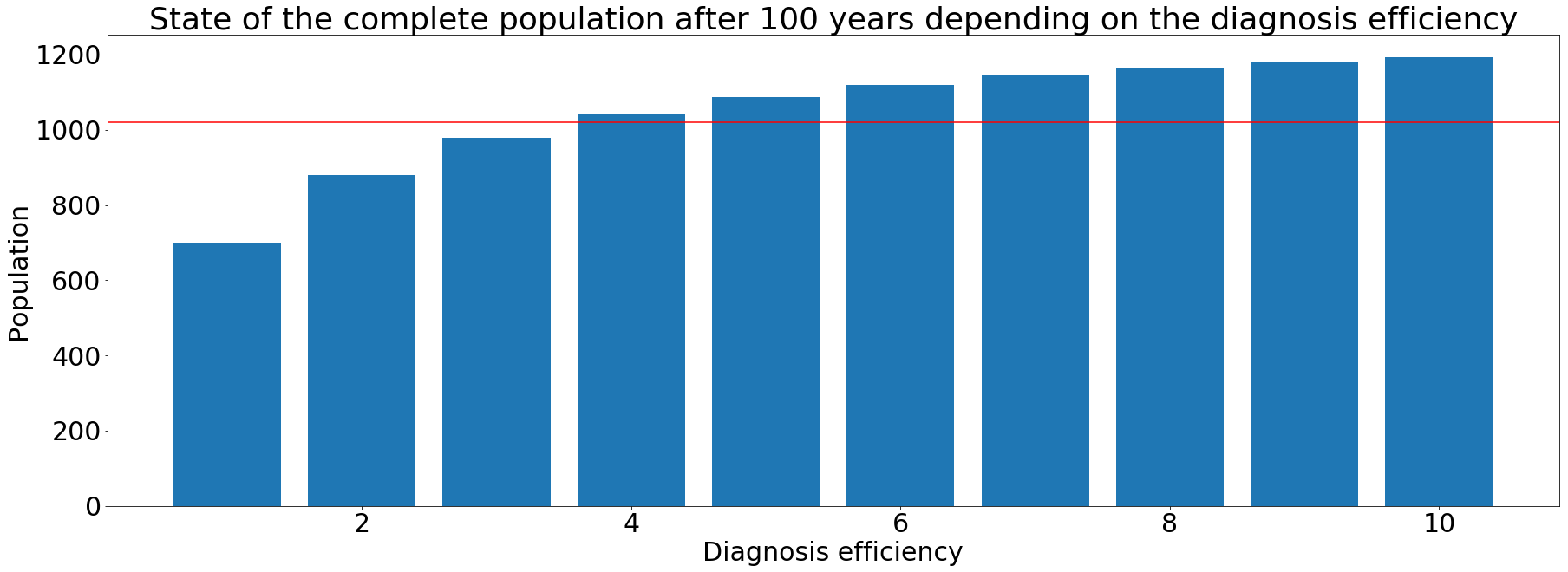
To help shape our design, we created an epidemiological model based on Abu-Raddad et al's paper. The model evaluates different cases of Tuberculosis dissemination and evolution throughout many years. Using the model, we saw by which disease state the population’s death rate is mostly affected. Then, we could concentrate our efforts on the diagnosis of that state. The increase in diagnosis efficiency showed promising results. Simulations showed that our diagnosis procedure helped decrease TB cases. The model is robust because it uses data from the world health organization and a complete scientists consortium. Though the model is complex; it includes multiple parameters. It has already allowed us to draw several conclusions. As the model improves and the parameters are refined, other conclusions can follow. Thus, our model helps understand critical aspects of the suggested design.
The engagement with the public, the communication and the sharing of our project was very important for us, in order to make known the stakes of the tuberculosis and its diagnosis, but also to make known the synthetic biology and the iGEM. Exchanges with specialists (biologists and doctors) has been crucial for the development of our kit to make it easy to use, cheap, effective, accurate and accessible. Our Human Practices and Public Engagement were very strong, fulfilling the silver medal criteria, and we also integrated our work on human practices into the design of the project as gold medal criteria.